Solutions
Video
Code
1. Consider the following blocks of code. Each one of them will cause an error when run. Taking each block one-by-one:
Predict which line of code will cause the error and why this line is problematic.
Run the block of code in a Jupyter notebook and see whether or not your were correct.
Modify the code to solve the problem and thus remove the error.
Block 1
You will need this file.
import numpy as np
wavenumber, transmittance = np.loadtxt("ir_data.csv", delimiter=",")
print(wavenumber, transmittance)Multiple assignment is performed but, by default, np.loadtxt only returns a single variable. Instead, the unpack keyword argument should be set to True in the call to np.loadtxt.
import numpy as np
wavenumber, transmittance = np.loadtxt("ir_data.csv", delimiter=",", unpack=True)
print(wavenumber, transmittance)Block 2
This block also contains a problem which, whilst it will not cause an error, will lead to the wrong result. See if you can spot and fix this problem in addition to removing the error.
You will need this file.
with open('elements.txt', 'a') as f:
lines = f.readlines()
for line in lines:
num_h += line.count('h')
print(f'There are {num_h:d} occurences of the letter H in the names of the 118 elements.')- The file is opened with
mode='a'(append), and should instead be opened withmode='r'(read). - The variable
num_his not defined before it is used with the+=operator. It should instead be initialised as equal to zero, and then updated in the loop. - Additionally, the method used to count the letter
Hwill only consider lower-case letters. Instead, both lower- and upper-case versions of the letter should be counted, or alternatively each line should be converted to lower-case.
Solution 1:
with open('elements.txt', 'r') as f:
lines = f.readlines()
num_h = 0
for line in lines:
num_h += line.count('h')
num_h += line.count('H')
print(f'There are {num_h:d} occurences of the letter H in the names of the 118 elements.')Solution 2:
with open('elements.txt', 'r') as f:
lines = f.readlines()
num_h = 0
for line in lines:
num_h += line.lower().count('h')
print(f'There are {num_h:d} occurences of the letter H in the names of the 118 elements.')2. The atomic coordinates of molecules are often stored xyz files. These have the following format:
NUMBER_OF_ATOMS
COMMENT
ATOM1_LABEL ATOM1_X ATOM1_Y ATOM1_Z
ATOM2_LABEL ATOM2_X ATOM2_Y ATOM2_Z
...
ATOMN_LABEL ATOMN_X ATOMN_Y ATOMN_ZThe number of atoms is an int, the comment line is a str which can optionally be blank, the atom labels are str (e.g. 'H', or 'H2'), and the \(x\), \(y\), and \(z\) coordinates are all float with units of Angstrom.
a. Using np.loadtxt, read this xyz file for hydrogen peroxide (\(\mathrm{H_2O_2}\)) and store its atom labels and coordinates in two separate NumPy arrays.
The labels and coordinates can be read in separately using two calls to np.loadtxt:
import numpy as np
labels = np.loadtxt('h2o2.xyz', skiprows=2, usecols=0, dtype=str)
coords = np.loadtxt('h2o2.xyz', skiprows=2, usecols=(1, 2, 3))
print(coords)
print(labels)where in both cases skiprows=2 has been used to ignore the first two lines of the file, since these have a different format to the remaining lines. In the first call, usecols=0 specifies that only the first column of data is extracted, and dtype=str specifies that the data should be read as strings. In teh second call, usecols=(1, 2, 3) is specified in order to only extract the coordinate columns, while dtype is not given since by default numopy expects to read a float.
The labels used in this xyz file are shown in the figure below.
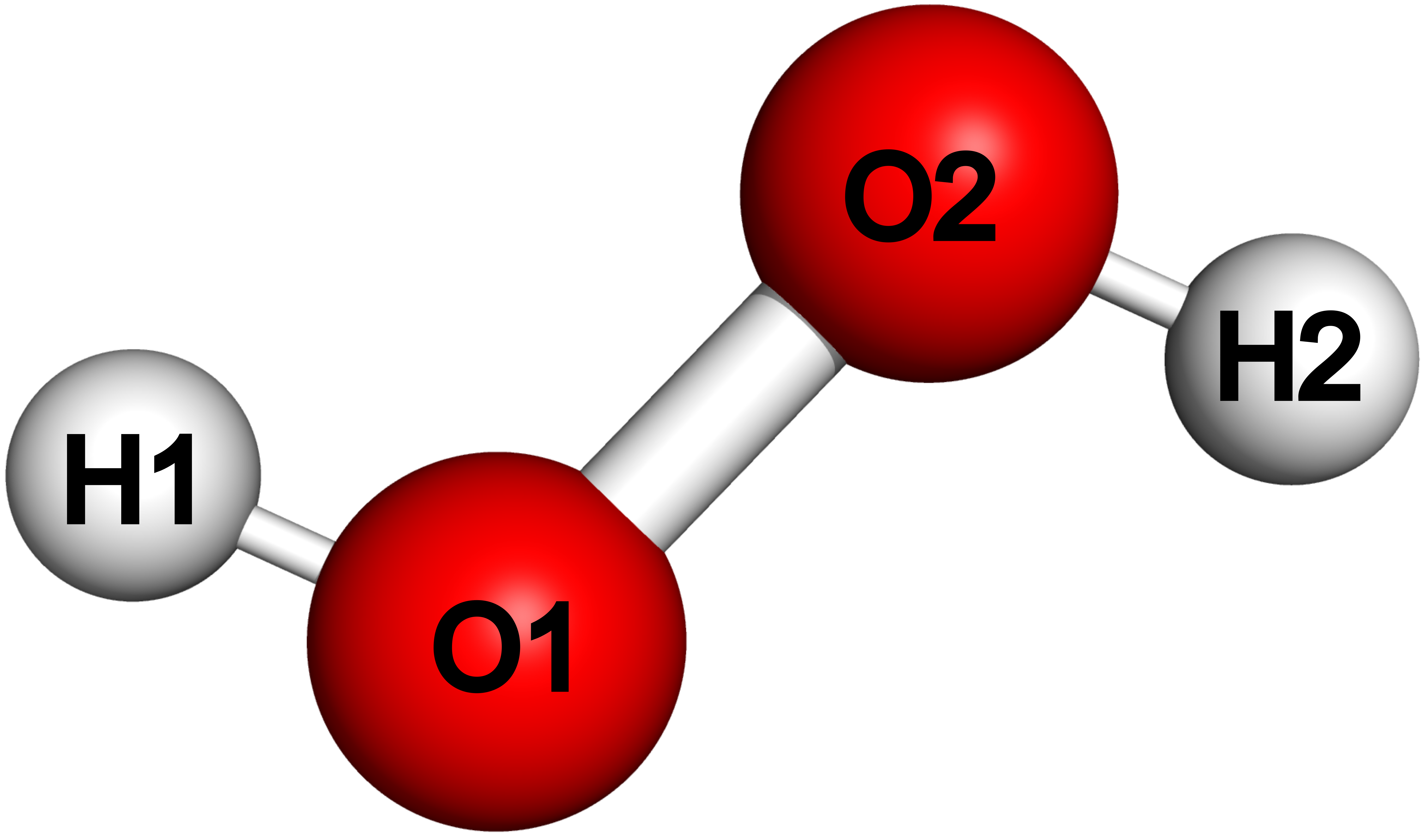
b. The distance between two atoms \(\mathrm{A}\) and \(\mathrm{B}\) is given by
\[\begin{equation*} d_\mathrm{AB} = \sqrt{\left(x_\mathrm{B}-x_\mathrm{A}\right)^2 + \left(y_\mathrm{B}-y_\mathrm{A}\right)^2 + \left(z_\mathrm{B}-z_\mathrm{A}\right)^2} \end{equation*}\]
where, for example, \(x_\mathrm{B}\) is the \(x\) coordinate of atom \(\mathrm{B}\).
i. Write a function that takes two NumPy arrays as arguments - each containing the \(x\), \(y\), and \(z\) coordinates of an atom - and returns the distance between them.
def calc_distance(atom_1, atom_2):
'''
Calculates the distance between two atoms.
Arguments
---------
atom_1: np.ndarray
x, y, and z coordinates of atom 1 in Angstrom
atom_2: np.ndarray
x, y, and z coordinates of atom 2 in Angstrom
Returns
-------
float
Distance between two atoms in Angstrom
'''
dist = (atom_2[0] - atom_1[0])**2 + (atom_2[1] - atom_1[1])**2 + (atom_2[2] - atom_1[2])**2
dist = np.sqrt(dist)
return distOur function computes the equation for a given pair of np.array objects. Remember to include a docstring with your function!
If you remember that we can also calculate the distance between two atoms using vector operations, you could instead write the function much more concisely as
def calc_distance(atom_1, atom_2):
'''
Calculates the distance between two atoms.
Arguments
---------
atom_1: np.ndarray
x, y, and z coordinates of atom 1 in Angstrom
atom_2: np.ndarray
x, y, and z coordinates of atom 2 in Angstrom
Returns
-------
float
Distance between two atoms in Angstrom
'''
dist = np.sqrt(np.sum((atom_2 - atom_1)**2))
return distii. Using your function, calculate the \(\mathrm{O}_1-\mathrm{O}_2\) bond length, and the average \(\mathrm{O}-\mathrm{H}\) bond length.
# Oxygen-Oxygen bond distance
O1O2_distance = calc_distance(coords[0], coords[1])
# Oxygen-Hydrogen bond distances
O1H1_distance = calc_distance(coords[0], coords[2])
O2H2_distance = calc_distance(coords[1], coords[3])
OH_avg_distance = (O1H1_distance + O2H2_distance) / 2
print(f'The oxygen-oxygen bond distance is {O1O2_distance:.3g} Å')
print(f'The average oxygen-hydrogen bond distance is {OH_avg_distance:.3g} Å')In the first call to calc_distance, we’ve indexed the coords array from part a to select only the rows which correspond to the oxygen atoms in hydrogen peroxide. We can figure out that the first and second row are the oxygen atoms by looking at the labels array which is printed in part a - this will be in the same order as coords since they were read from the same file.
The same is true for the further calls to calc_distance where we calculate the two \(\mathrm{OH}\) bond lengths. We then take a simple average of the two and print all the information to screen.
c. We can define the vector \(\overrightarrow{\mathrm{AB}}\) between atoms \(\mathrm{A}\) and \(\mathrm{B}\) as
\[\begin{equation*} \overrightarrow{\mathrm{AB}} = \begin{bmatrix} x_\mathrm{B}-x_\mathrm{A}\\ y_\mathrm{B}-y_\mathrm{A}\\ z_\mathrm{B}-z_\mathrm{A} \end{bmatrix} \end{equation*}\]
A vector is a mathematical quantity which has both direction and magnitude.
We denote vectors using an arrow over the top of their mathematical symbol, and write them in full as either a column or a row containg a set of coordinates which describe the vector’s magnitude (length) and direction.
c.
Calculate the vector \(\overrightarrow{\mathrm{O_1O_2}}\) which connects the two oxygen atoms \(\mathrm{O_1}\) and \(\mathrm{O_2}\), and store it in a NumPy array.
oo_vector = np.array([
coords[0, 0] - coords[1, 0],
coords[0, 1] - coords[1, 1],
coords[0, 2] - coords[1, 2]
])
print(oo_vector)[0. 0. 1.48]Again, we use the first and second rows of coords which store the \(x\), \(y\), and \(z\) coordinates of \(\mathrm{O_1}\) and \(\mathrm{O_2}\), respectively.
d. Using np.linalg.norm, compute the vector norm of \(\overrightarrow{\mathrm{O_1O_2}}\) from part c. Compare this to your answer in part b, what do you think the vector norm is?
print(f'The vector norm of the O1O2 vector is {np.linalg.norm(oo_vector):.3g} Å')1.48 ÅNotice that this is exactly the same result as in b - the vector norm is the length (or magnitude) of a vector:
\[\begin{equation*} \left|\overrightarrow{\mathrm{AB}}\right| = \sqrt{\left(\overrightarrow{\mathrm{AB}}_x\right)^2 + \left(\overrightarrow{\mathrm{AB}}_y\right)^2 + \left(\overrightarrow{\mathrm{AB}_z}\right)^2} \end{equation*}\]
we square each element of the vector, sum them, and take the square root the resulting number.
e. The angle \(\theta\) between two bonds can be obtained by constructing vectors between the three atoms involved and then calculating the angle between those vectors.
\[\begin{equation*} \theta = \cos^{-1}\left(\frac{\overrightarrow{\mathrm{AB}}\cdot\overrightarrow{\mathrm{BC}}}{d_\mathrm{AB}\, d_\mathrm{BC}}\right) \end{equation*}\]
where the symbol \(\cdot\) implies the dot product of the two vectors is taken. This is defined as
\[\begin{equation*} \overrightarrow{\mathrm{D}}\cdot\overrightarrow{\mathrm{E}} = x_\mathrm{D} x_\mathrm{E} + y_\mathrm{D} y_\mathrm{E} + z_\mathrm{D} z_\mathrm{E} \end{equation*}\]
i. Write a function that takes two NumPy arrays as arguments - each containing a vector - and returns the angle between them.
def vector_angle(vector_1, vector_2):
'''
Computes the angle between two vectors
Arguments
---------
vector_1: np.ndarray
First vector
vector_2: np.ndarray
Second vector
Returns
-------
float
Angle between vectors in degrees
'''
length_1 = np.linalg.norm(vector_1)
length_2 = np.linalg.norm(vector_2)
numerator = np.dot(vector_1, vector_2)
denominator = length_1 * length_2
angle = np.arccos(numerator/denominator)
angle = np.rad2deg(angle)
angle = min(angle, 180-angle)
return angleNotice we’ve added the line angle = min(angle, 180-angle) - this is important since the output of np.arccos sometimes gives us the outside angle between two vectors. When discussing bond angles we always use the inside angle, so we calculate the smaller of angle and 180 - angle.
ii. Using your function, calculate the average \(\mathrm{O\hat{O}H}\) bond angle in hydrogen peroxide.
# Calculate each OH bond vector
o1h1_vector = np.array([
coords[2, 0] - coords[0, 0],
coords[2, 1] - coords[0, 1],
coords[2, 2] - coords[0, 2]
])
o2h2_vector = np.array([
coords[3, 0] - coords[1, 0],
coords[3, 1] - coords[1, 1],
coords[3, 2] - coords[1, 2]
])
# H1-O1-O2 angle
h1o1o2_angle = vector_angle(oo_vector, o1h1_vector)
# H2-O2-O1 angle
h2o2o1_angle = vector_angle(oo_vector, o2h2_vector)
average_angle = (h1o1o2_angle + h2o2o1_angle) / 2
print(f'The average OOH angle is {average_angle:.3g}°')The average OOH angle is 70.5°